16 November 2015
Mysteries of Bony Fish Genome Evolution
Mistakes Propelling Evolution
As the 20th-century novelist Joseph Conrad famously wrote, “It’s only those who do nothing that make no mistakes, I suppose,” and Nature is very busy, so she makes lots of them. But as a genius, she can use them to advantage. Take for example whole genome duplication — an error in DNA replication, or mating between different species, that doubles the number of chromosomes, leading to a duplication of the vast majority of genes. Such grand mistakes turn out to be among the major forces accelerating evolution.
Organisms with additional sets of genes can accumulate and test mutations much faster and with less selection pressure than organisms with just one set of genes. One of the copies of a gene can maintain normal functioning of the cells even if the other copy mutates to become harmful or useless. Other possible alternatives are when one of the genes acquires a completely new function, or both genes start to specialise, each taking over a certain part of the ancestral function.
Let’s Talk Fish
Redundancy is common in nature, for example many human genes exist in several copies. However, excessive redundancy can interfere with efficiency. Therefore, each whole genome duplication event is followed by the loss of duplicate genes.
A new collaborative paper, published in Proceedings of the National Academy of Sciences (PNAS) by scientists from the Okinawa Institute of Science and Technology Graduate University (OIST), the University of the Ryukyus, Tohoku University, and Nihon University, proposes a two-phase mathematical model describing gene loss patterns over two differing time scales after a whole genome duplication.
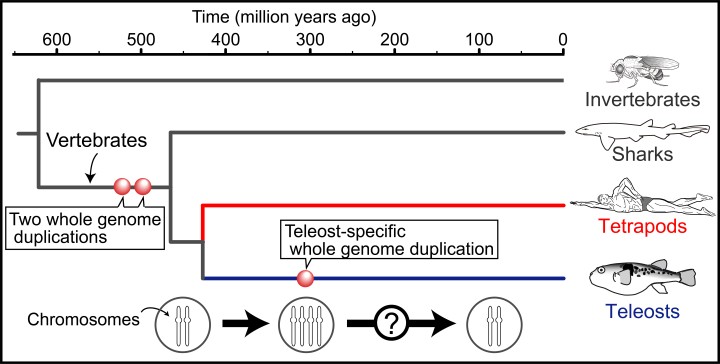
The research, started by Dr Jun Inoue, now a staff scientist in the OIST Mathematical Biology Unit, Assistant Prof. Yukuto Sato, now at Tohoku University, and Prof. Mutsumi Nishida, now Vice President of the University of the Ryukyus, while all three were working together at the University of Tokyo, focused on teleost fishes, the largest group of bony fishes (Fig. 1). These fishes underwent their own specific whole genome duplication approximately 300 million years ago. “By contrast, the last whole genome duplication in the human lineage happened about half a billion years ago, and it is extremely difficult to trace,” explains Prof. Robert Sinclair, the head of the Mathematical Biology Unit.
The work required the development of both new computational and also mathematical tools, each one adapted to the previously determined evolutionary history of the species involved. These tools, developed by the research group, can be applied to other cases of whole genome duplications in organisms of any kind, including humans.
Genome Bulk Editing
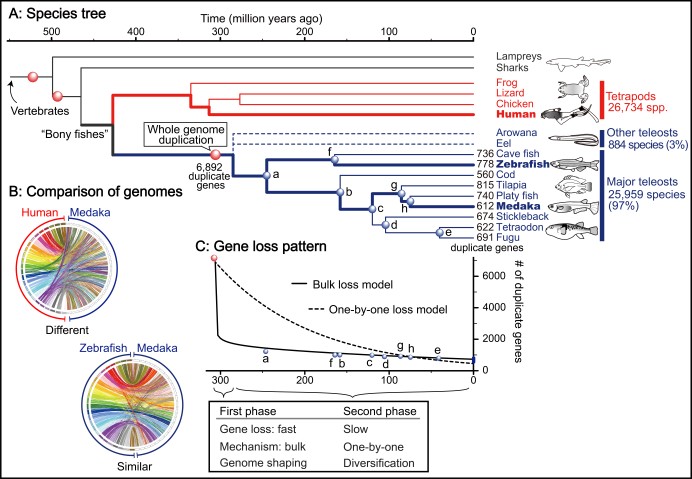
Comparison of the genomes of zebrafish and the common Japanese fish medaka — two distantly related species with 250 million years of independent evolution (Fig. 2A) — shows that they are very similar (Fig. 2B). “Their shapes, habitats, and reproductive patterns are very different, which suggests that the basic structure of the teleost genome was established before the major diversification of teleost species,” says Dr Inoue. Genome analysis of seven other well-studied fishes supports this conclusion.
The results of this study suggest that approximately 80% of the duplicate genes were lost in the first 60 million years after the whole genome duplication event (Fig. 2C). Considering that the first vertebrates appeared on Earth about 500 million years ago (Fig. 2A), 60 million years is a very short time. Dr Inoue states that it is possible that genome reduction happened even faster. “We are waiting for the eel genome to be fully decoded to check this hypothesis,” he says. Eels and their relatives are one of the first groups separated from the majority of teleost fishes after the teleost-specific whole genome duplication. Comparison of eels with the other teleosts will eventually shed even more light on the evolution of all fishes.
The idea of loss of redundant genes is not new; however, an important new result is that “we found evidence that genes are disposed of rapidly and in bulk after the whole genome duplication, and it leads to a rapid reshaping of the genome,” says Prof. Sinclair. In the first phase, clusters of adjacent genes or even large chromosomal segments may have been deleted if they were useless or problematic. Interestingly, the diversification of many major lineages of living teleosts did not occur in this rapid phase (Fig. 2A).
The second phase is characterised by slower gene loss (Fig. 2C). The scientists suggest that some paired genes are retained if each copy becomes essential. Other duplicate genes continue to be lost, but mostly one-by-one. This process continues to this day. Evolution never stops!
The methods of genome analysis developed for this study will pave the road to a better understanding of evolution, including our own.