A representation of the genome assembly method
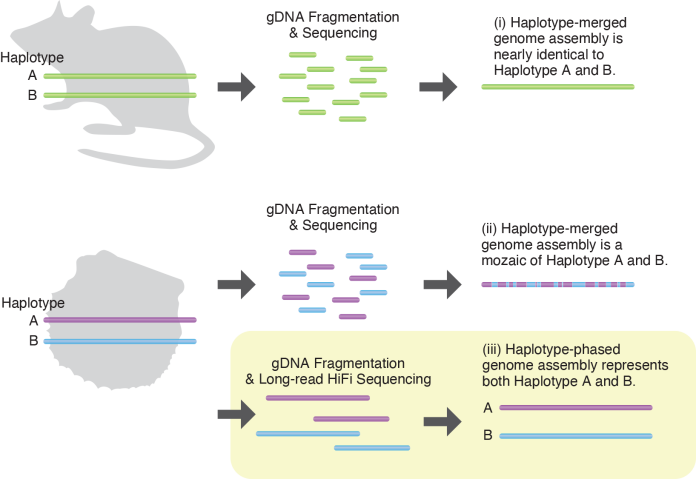
The genetic information necessary for an organism to maintain its vital activities is called a genome. In genome decoding, DNA is extracted from individual cells, fragmented, and analyzed. The DNA sequence fragments are then reconstructed to obtain a genome assembly. Animals that reproduce sexually inherit one set of genome from the mother and one from the father. A set of genomic information derived from one parent is called a haplotype. (i) In experimental organisms with established strains or species with small genetic diversity, an individual possesses two sets of nearly identical genomes. Thus, the haplotype-merged genome assembly will be similar to both the two sets of genomes of the original individual. (ii) In organisms with high genetic diversity, such as wild animals, there are large differences in DNA sequences among haplotypes. Using conventional methods results in a genome assembly with a mixture of two haplotypes. It can lose genomic information. (iii) In this study, longer and more accurate DNA sequences were obtained by using the latest sequencer. The two haplotypes were reconstructed separately.
The genetic information necessary for an organism to maintain its vital activities is called a genome. In genome decoding, DNA is extracted from individual cells, fragmented, and analyzed. The DNA sequence fragments are then reconstructed to obtain a genome assembly. Animals that reproduce sexually inherit one set of genome from the mother and one from the father. A set of genomic information derived from one parent is called a haplotype. (i) In experimental organisms with established strains or species with small genetic diversity, an individual possesses two sets of nearly identical genomes. Thus, the haplotype-merged genome assembly will be similar to both the two sets of genomes of the original individual. (ii) In organisms with high genetic diversity, such as wild animals, there are large differences in DNA sequences among haplotypes. Using conventional methods results in a genome assembly with a mixture of two haplotypes. It can lose genomic information. (iii) In this study, longer and more accurate DNA sequences were obtained by using the latest sequencer. The two haplotypes were reconstructed separately.
Read the associated press release: Molecular fingerprint behind beautiful pearls revealed
Copyright OIST (Okinawa Institute of Science and Technology Graduate University, 沖縄科学技術大学院大学). Creative Commons Attribution 4.0 International License (CC BY 4.0).














